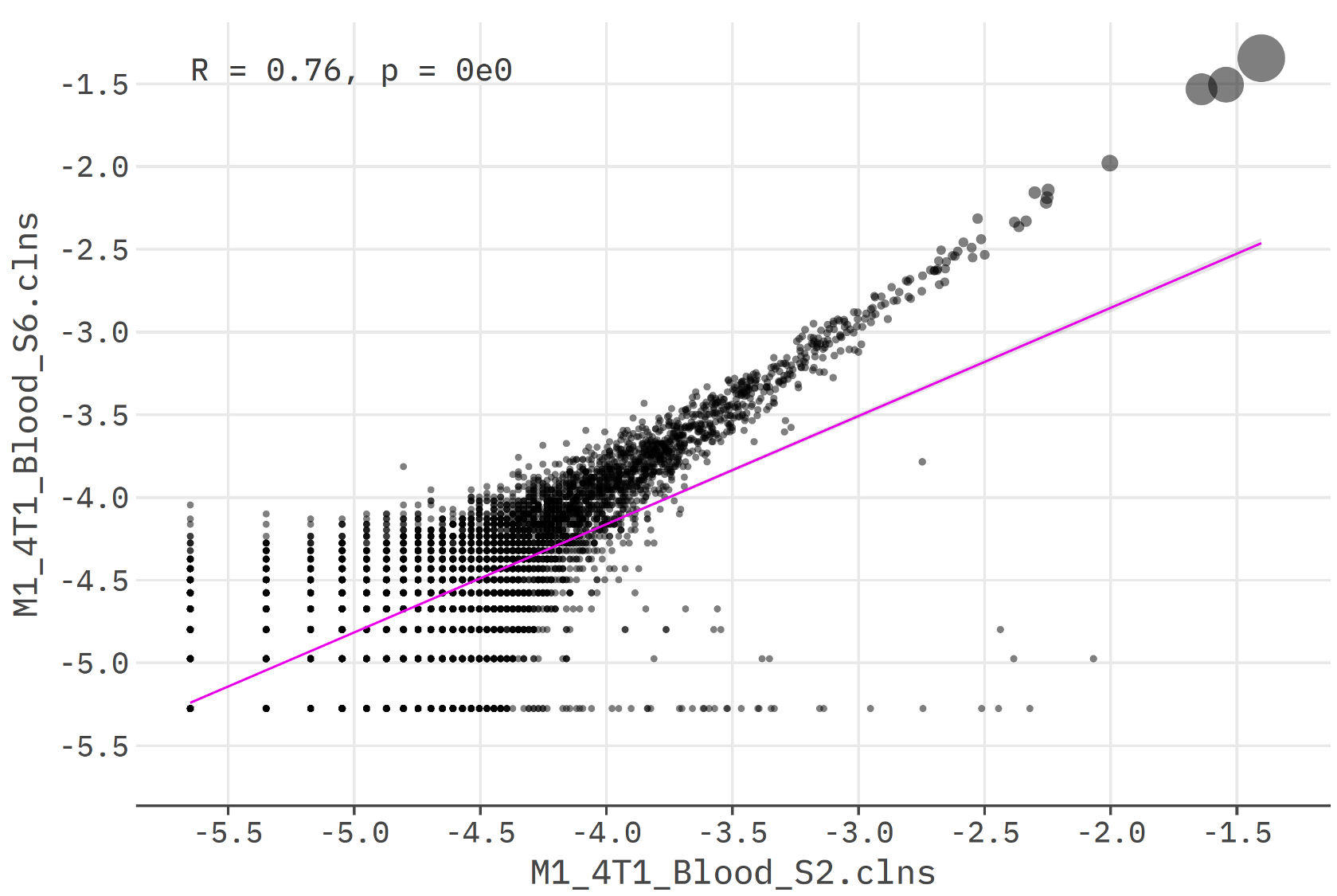
mixcr overlapScatterPlot
mixcr overlapScatterPlot creates a scatter-plot for clone frequencies overlap between two samples.
Command line options
mixcr overlapScatterPlot --downsampling (<type>|none) [--chains <chain>[,<chain>...]]... [--only-productive] [--criteria <s>] [--method <method>] [--no-log] [--force-overwrite] [--no-warnings] [--verbose] [--help] cloneset_1.(clns|clna) cloneset_2.(clns|clna) output.(pdf|eps|svg|png|jpeg)
The command takes .clna or .clns file as input and produces one of the following graphical formats depending on the extension of output file: .pdf, .eps, .png, .svg and .jpeg
--downsampling (<type>|none)- Choose downsampling applied to normalize the clonesets. Possible values:
count-[reads|TAG]-[auto|min|fixed][-<number>],top-[reads|TAG]-[<number>],cumtop-[reads|TAG]-[percent],none --chains <chain>[,<chain>...]- Chains to export.
--only-productive- Filter out-of-frame sequences and sequences with stop-codons.
--criteria <s>- Overlap criteria. Defines the rules to treat clones as equal. Default:
CDR3|AA|V|J(For two clones to me equal they must shareCDR3amino acid sequence, V and J genes) --method <method>- Correlation method to use. Possible value: Pearson, Kendall, Spearman. Default: Pearson
--no-log- Do not apply log10 to clonotype frequencies.
-f, --force-overwrite- Force overwrite of output file(s).
-nw, --no-warnings- Suppress all warning messages.
--verbose- Verbose messages.
-h, --help- Show this help message and exit.
Example:
```shell
mixcr overlapScatterPlot \ --downsampling none \ --chains IGH \ results/M1_4T1_Blood_S2.clns results/M1_4T1_Blood_S6.clns \ M1_4T1_Blood.overlap.pdf ```